|
The aims of our study were 1) to compare the performance of AmpliSeq and SureSelect and 2) to develop an optimized protocol for variant calling for WES on Ion Proton platform. Studies comparing the only two library preparation methods, namely, AmpliSeq and SureSelect for Ion Proton, are, however, so far missing. A recent in-detail evaluation of AmpliSeq WES using NA12878 reference calls ( Damiati et al., 2016) highlighted the limitations of PCR-based target enrichment, provided the list of missed target regions and a filtering strategy to reduce the number of false positives (FPs). They found that AmpliSeq on Ion platforms is a faster method with high throughput but faces problem in complex genomic regions. Previous studies have compared AmpliSeq on Ion platforms with various kits available for Illumina platforms for WES ( Loman et al., 2012 Boland et al., 2013 Samorodnitsky et al., 2015). Studies comparing these two WES library preparation methods are so far missing. On the contrary, for Ion Proton, only two library preparation methods exist. Most of the library preparation methods are designed for Illumina platforms, and several articles compared the performances of these methods ( Clark et al., 2011 Lelieveld et al., 2015 Shigemizu et al., 2015). Library preparation is a critical step prior to enrichment, as it includes the targeted probe-based capture or amplification of target regions from genomic DNA. The major steps in the NGS workflow are library preparation, sequencing, and data analysis. There are several NGS platforms available, but the largest share is of Illumina platforms followed by Ion Proton ( Goodwin et al., 2016). Although whole-exome sequencing (WES) targets less than 2% of the genome ( van Dijk et al., 2014), it is a cost-effective way to detect both common and rare variants in protein-coding regions. 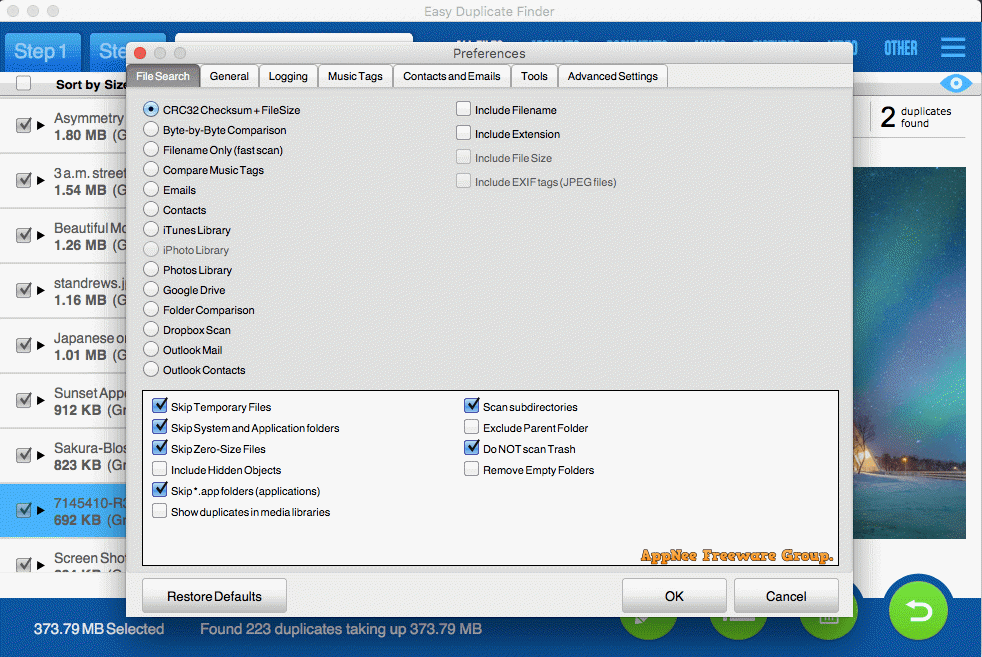
The use of whole-genome sequencing is yet limited due to its high cost. With the evolution of next-generation sequencing (NGS), base-by-base characterization of the human genome became possible.  
Genome-wide association studies (GWAS) using microarray-based genotypes in large-scale epidemiological studies played an essential role in the dissection of complex traits ( Kiezun et al., 2012). This improvement is highly relevant in research as well as clinical setting. Application of our variant calling pipeline decreased the number of false positive variants dramatically by 90% and resulted in positive predictive value of 97%. Both methods had a high concordance (>97%) with microarray genotypes and, when validating against NA12878, a sensitivity and positive predictive values of >93% and >80%, respectively. We used 12 in-house DNA samples with genome-wide and exome microarray data and a commercially available reference DNA (NA12878) for evaluation. Here, we systematically evaluate the performance of AmpliSeq and SureSelect and present an improved variant calling pipeline. Although of major interest, a comparison of the two methods is hitherto missing in the literature. For Ion Proton, there are only two exome library preparation methods available, AmpliSeq and SureSelect. Passware Windows Key Enterprise Edition 10.Library preparation for whole-exome sequencing is a critical step serving the enrichment of the regions of interest. 
QuickTime Pro 7.71.80.42 Lite + QuickTime Player PhotoInstrument 4.5.478 + MakeUpInstrument 4.4.444įlash Player 11.1.102.55 Projector StandAlone Hard Drive Inspector for Notebooks 3.93.421 NET Framework 1.1-4.0 for Windows XP SP3 x86 Microsoft Visual C++ 2005-2008-2010 x86/圆4Īdobe Flash Player 11.1.102.55 ActiveX & PluginĪdobe Shockwave Player 11.6.3.633 Full + Authorware Web Player Norton Internet Security 2012 19.1.1.3 Rus
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
